Metabolic flexibility and an unusual route for peptidoglycan muramic acid recycling in mycobacteria
Metabolic flexibility and an unusual route for peptidoglycan muramic acid recycling in mycobacteria
Stravoravdis, S.; Carnahan, B.; Gordon, R. A.; Wodzanowski, K.; Havaleshko, K.; Fils-Aime, E.; Putnik, R.; Hyland, S.; Grimes, C. L.; Siegrist, M. S.
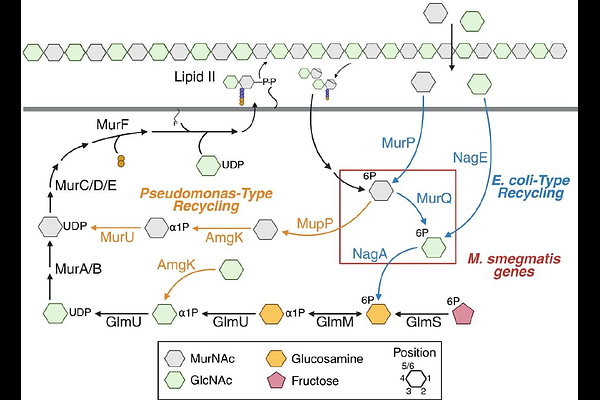
AbstractDe novo biosynthesis of cell wall peptidoglycan is essential for bacterial viability under many growth conditions and is a well-validated antibiotic target. Although generally not essential for bacterial fitness under standard laboratory growth conditions, peptidoglycan recycling can aid bacterial survival under host or antibiotic stress. Peptidoglycan consists of alternating sugars N-acetylmuramic acid (MurNAc) and N-acetylglucosamine (GlcNAc) cross-linked by peptides. Recycling of these sugars can proceed via GlcNAc and glucosamine intermediates (Escherichia coli-type) or, in the case of MurNAc, bypass these intermediates altogether (Pseudomonas-type). We serendipitously discovered that the pathogen Mycobacterium tuberculosis and model organism M. smegmatis assimilate 2-modified MurNAc probes into their peptidoglycan despite lacking the Pseudomonas-type machinery that is normally required for incorporation of these molecules. Our data suggest that unmodified and 2-modified MurNAc incorporate into M. smegmatis peptidoglycan via multiple pathways, the former preferentially via an E. coli-type route and the latter preferentially via a non-E. coli, non-Pseudomonas-type route with GlcNAc but not glucosamine intermediates. These findings reveal metabolic flexibility in mycobacterial cell wall recycling that encompasses a previously undescribed pathway.