SARS-CoV-2 Defective Viral Genomes from Distinct Genomic Regions Drive Divergent Interferon Responses
SARS-CoV-2 Defective Viral Genomes from Distinct Genomic Regions Drive Divergent Interferon Responses
Brennan, J. W.; Spandau, S.; Wang, X.; Aull, H.; Connor, S.; Pryhuber, G.; Mariani, T. J.; Serra-Moreno, R.; Sun, Y.
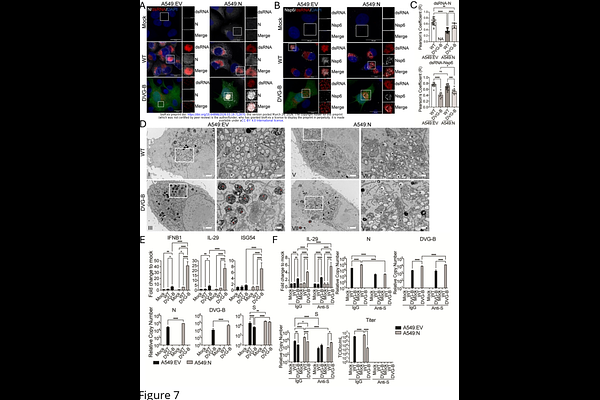
AbstractDefective viral genomes (DVGs) are naturally generated during genomic replication of many RNA viruses. When produced early in infection or supplemented at the onset of infection, DVGs can attenuate viral pathogenesis by stimulating IFN responses and antagonizing wild type (WT) virus replication, highlighting their potential as antiviral therapeutics. However, during natural infection DVGs can exert both antiviral and proviral effects depending on their generation kinetics, species, and abundance, underscoring the need to better understand their roles in viral pathogenesis. Coronaviruses (CoVs) remain a major global health threat and ubiquitously generate DVGs, yet DVGs roles during CoV infection are largely unknown. Using SARS-CoV-2 as a model, we previously identified DVGs in vitro and in patient samples and discovered two major genomic hotspots (A and B) for their generation. Here, we first showed that overall DVG abundance tended to positively correlate with COVID-19 severity, with approximately 40% of DVGs originating from a specific genomic region designated hotspot B. Analysis of a publicly available single-cell RNA-seq datasets revealed that DVGs from hotspot B, but not hotspot A, were associated with elevated IFN responses, suggesting that DVGs derived from different genomic regions vary in their ability to stimulate innate immunity. To test this directly, we constructed two representative DVGs corresponding to hotspots A and B. Both DVGs suppressed the replication of co-infecting WT virus; however, only DVG-B induced robust IFN responses, exceeding those triggered by WT virus alone. This was further confirmed in human precision-cut lung slices. Mechanistically, DVG-B-derived dsRNA exhibited a distinct subcellular distribution compared with WT virus. Complementation with the nucleocapsid (N) partially restored dsRNA organization but did not alter the IFN response. Together, our findings demonstrate that DVGs arising from distinct genomic hotspots differentially regulate IFN responses, potentially contributing to varied pathogenic outcomes during SARS-CoV-2 infection.