Identification and characterization of a poly(ϵ-caprolactone)-degrading enzyme with a unique sequence profile from the marine bacterium Alloalcanivorax gelatiniphagus
Identification and characterization of a poly(ϵ-caprolactone)-degrading enzyme with a unique sequence profile from the marine bacterium Alloalcanivorax gelatiniphagus
Kusumoto, H.; Hachisuka, S.-i.; Iseki, K.; Kikukawa, H.; Matsumoto, K.
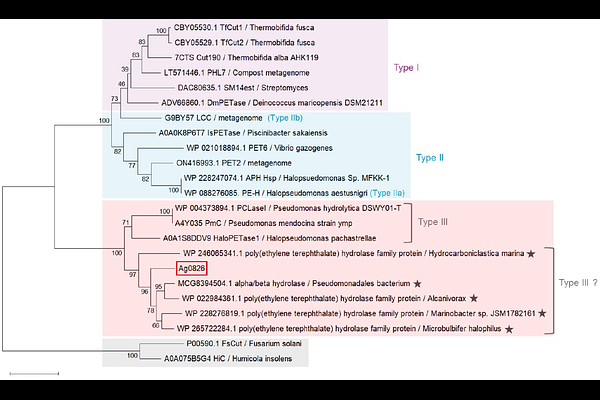
AbstractPoly({epsilon}-caprolactone) (PCL) is a well-known biodegradable polyester and is among the few polyesters susceptible to degradation in marine environments; however, marine-derived PCL-degrading enzymes remain poorly characterized. Here, we searched for PCL-degrading enzymes from the marine bacterium Alloacanivorax gelatiniphagus JCM 18425 using a genome-based approach. Five candidate genes were predicted, and one encoded protein, designated Ag0826, was identified as a PCL depolymerase. Recombinant Ag0826 was expressed, purified, and biochemically characterized. The enzyme exhibited optimal activity at 35-40{degrees}C and pH 8.0, although it showed limited thermal stability. Substrate specificity was compared with that of leaf-branch compost cutinase (LCC), a well-characterized poly(ethylene terephthalate) (PET) hydrolase, using various polyesters. Both enzymes exhibited largely overlapping substrate ranges with respect to the presence or absence of monomer conversion activity across the tested substrates. Ag0826 slightly degraded PET to terephthalic acid, indicating potential PET-hydrolyzing activity; its conversion rate, however, was substantially lower than that of LCC, suggesting that Ag0826 exhibits a substrate preference differing from LCC. Phylogenetic analysis based on amino acid sequences revealed that Ag0826 formed a separate clade from LCC and IsPETase (from Ideonella sakaiensis). At a broader level, Ag0826 was positioned near HaloPETase1 (from Halopseudomonas pachastrellae), which has been proposed as a Type III PET hydrolase; in contrast, residues corresponding to the substrate-binding subsites were similar but not identical between the two enzymes. These results suggest that Ag0826 broadly belongs to the group of known PET hydrolases, yet it exhibits a partially distinct sequence profile even within this enzyme family.