Spatial transcriptomics identifies dysregulated programs across neural and non-neural tissues in spinal muscular atrophy
Spatial transcriptomics identifies dysregulated programs across neural and non-neural tissues in spinal muscular atrophy
Rietz, A.; Kumari, L.; Androphy, E. J.
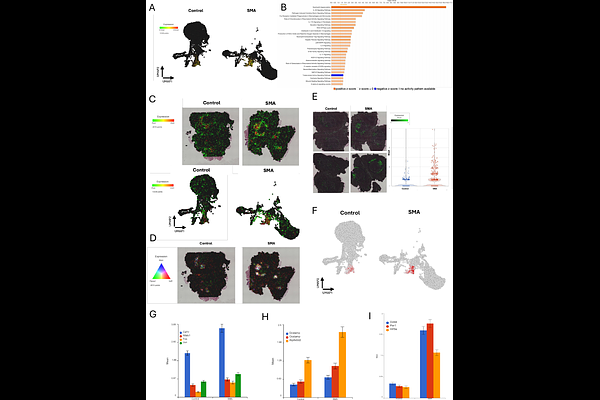
AbstractSpinal muscular atrophy (SMA) is caused by insufficient levels of the survival motor neuron (SMN) protein and clinically manifests as profound weakness due to motor neuron degeneration. While recent evidence suggests it is a multisystem disorder, the pathological programs in neural and peripheral tissues are poorly understood. We applied spatial transcriptomics to cross-sections from the entire lumbar region of pre-symptomatic SMA and control mice to define early, tissue-wide transcriptional consequences of SMN deficiency. This approach enabled spatially preserved analysis of spinal cord, dorsal root ganglia, muscle, bone, cartilage, bone marrow, adipose, and connective tissues within their native anatomical context. Within the SMA spinal cord, motor neuron associated regions and ventral interneurons exhibited upregulation of neurofilaments, tubulin isoforms, and microtubule transport machinery. Neurofilament changes were largely restricted to motor neurons, and tubulin dysregulation extended broadly across ventral regions. Multiple tissues displayed collagen and extracellular matrix gene dysregulation at post-natal day 4. Skeletal muscle demonstrated fiber type-specific stress responses, including induction of atrophy-associated genes, while bone displayed an osteoclast-dominant transcriptional signature, consistent with accelerated resorption and a pro-osteoporotic state. Bone marrow transcriptomes indicated activation of neutrophil degranulation, innate immune signaling, and osteoclastogenic pathways, identifying bone marrow as an active inflammatory and skeletal regulatory niche in SMA. Adipose tissue exhibited extracellular matrix dysregulation, complement activation, profibrotic TGF-{beta} signaling, and stress-induced lipolysis. These findings reveal that SMN deficiency drives early transcriptional reprogramming across multiple tissues well before motor neuron loss and identify non-neuronal pathological programs that offer therapeutic targets for improving long-term outcomes in SMA.