Bacterial Signatures and Community Structure in the Phyllosphere of Eugenia uniflora: Developmental Dynamics and Core Microbiome in Myrtaceae
Bacterial Signatures and Community Structure in the Phyllosphere of Eugenia uniflora: Developmental Dynamics and Core Microbiome in Myrtaceae
Cadavid Sanchez, I. C.; Esquen, D.; Margis, R.; Guzman Escudero, F. L.
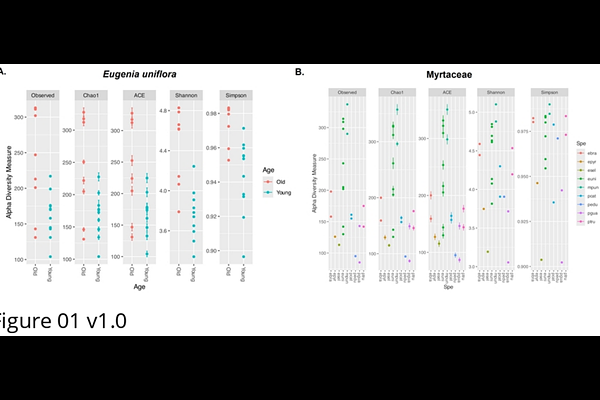
AbstractPlants recruit microorganisms to form mutually beneficial associations that enhance their health, productivity, and resilience. The composition of the plant microbiome is shaped by factors such as host species, developmental stage, genotype, and tissue type, with microbial recruitment mediated by plant exudates and secondary metabolites. Eugenia uniflora, a Myrtaceae species native to Brazils Atlantic Forest, produces pharmacologically relevant secondary metabolites and holds ecological and economic value. However, little is known about its associated microbiome, particularly from a metagenomic perspective. In this study, we investigated the phyllosphere bacterial communities, both epiphytic and endophytic, of E. uniflora across two developmental stages (young and mature trees). We also examined the core microbiome shared between E. uniflora and other Myrtaceae genera to better understand microbial diversity and structure within this family. Amplicon sequencing of the V3-V4 region of the 16S rRNA gene was conducted on 19 E. uniflora samples and 13 additional samples from three other Myrtaceae genera. In E. uniflora, we identified 1,456 bacterial ASVs representing 17 phyla, 115 families, and 171 genera. Alpha and beta diversity analyses revealed significant differences in bacterial community composition between developmental stages. Genera such as Massilia and Hymenobacter were more abundant in mature trees, while Aureimonas and Terriglobus were more common in young plants. Leaf microbiome functional potential shifted with plant age, with older leaves favoring secondary metabolite production and younger leaves emphasizing microbial interactions and defense. A total of 16 genera formed the Myrtaceae core microbiome, with five, Methylobacterium-Methylorubrum, Hymenobacter, Sphingomonas, Bdellovibrio, and Terriglobus, present in 100% of samples. Notably, ~0.7% of the bacterial diversity remained poorly classified, highlighting the underexplored nature of Myrtaceae-associated microbiomes and their potential for bioprospecting.