A Multi-Objective Scoring (MOS) Framework for Detecting Cross-Modal Spatial Similarity: Conceptual and Direct Formulations
A Multi-Objective Scoring (MOS) Framework for Detecting Cross-Modal Spatial Similarity: Conceptual and Direct Formulations
Jarrahi, A.; Jones, A. R.; Tang, W.; Qi, H.; Crouch, A. C.
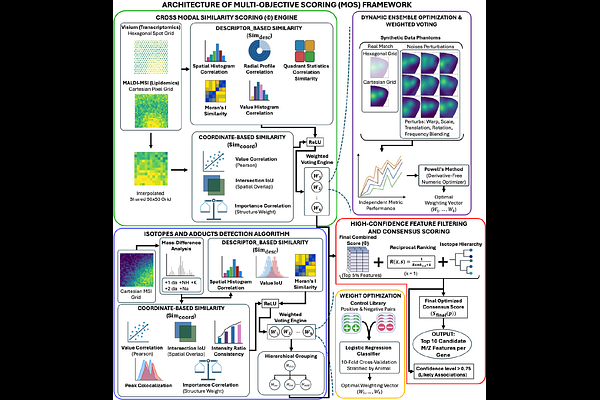
AbstractSpatial multi-omics methods allow researchers to study complex biological systems by integrating multiple molecular layers while preserving their spatial organization. However, integrating spatial transcriptomics and mass spectrometry imaging (MSI) remains challenging due to the differences between the two modalities, including sampling geometry, spatial resolution, signal scaling, and measurement principles. For example, 10x Genomics Visium captures transcriptomic data on a discrete hexagonal grid of spots; however, matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI) produces dense Cartesian pixel maps of molecular distributions. These differences make exact spatial co-registration difficult and limit the effectiveness of single-metric similarity approaches. Here, we introduce a Multi-Objective Scoring (MOS) framework designed to detect cross-modal spatial similarity without requiring exact pixel-to-pixel alignment. The MOS framework integrates multiple complementary spatial descriptors into a unified similarity score, including coordinate-based metrics (value correlation, importance-based Intersection-over-Union (IoU), and importance-map correlation) and descriptor-based metrics that capture higher-order spatial organization, such as spatial histograms, radial profiles, quadrant statistics, and Moran's I spatial autocorrelation. These metrics are combined through a weighted ensemble model. This model calculates the weights using synthetic spatial datasets that simulate realistic tissue geometry, sampling differences, and spatial distortions. The framework was applied to a spatial multi-omics dataset from murine brain tissue, integrating spatial transcriptomics with MALDI-MSI lipidomics across young and aged control and Alzheimer's disease (AD) models. Synthetic data validation results demonstrated strong pattern-matching performance (96.14% accuracy), and application to experimental data identified several MSI analyte features whose spatial distributions closely matched transcriptomic patterns. In particular, strong and reproducible associations were observed between myelin-related genes (Mbp and Plp1) and multiple analyte features enriched in white matter regions. Overall, whether applied conceptually or directly, the MOS framework provides a strong strategy for cross-modal spatial integration and offers a scalable tool for discovering spatial relationships across diverse multi-omics datasets and facilitating hypothesis generation.