Saliva Metabolomics Reveals Distinct Metabolic Signatures in Patients with Chronic Obstructive Pulmonary Disease: A GC-MS-based approach.
Saliva Metabolomics Reveals Distinct Metabolic Signatures in Patients with Chronic Obstructive Pulmonary Disease: A GC-MS-based approach.
Singh, R.; Ghosh, S.; Yadav, N.; Mandal, A. K.
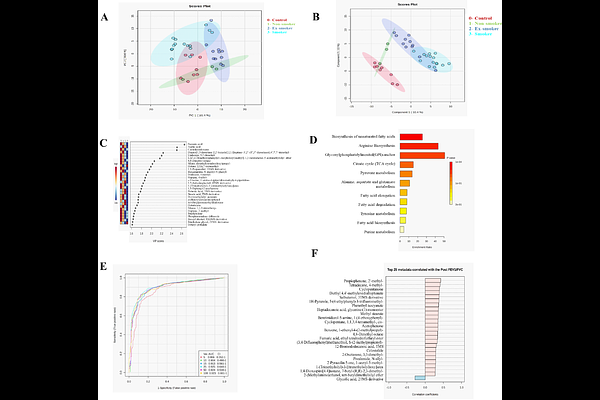
AbstractChronic obstructive pulmonary disease (COPD), a chronic lung disease, involves complex metabolic disturbances that remain poorly characterized using non-invasive matrices. The metabolic alterations associated with cigarette smoke (CS), one of the major drivers of disease progression in COPD patients, have not been explored in detail. This study primarily aimed to investigate the metabolic signatures in COPD patients categorized into smoker (n=15), ex-smoker (n=11), and non-smoker (n=3) subgroups. Utilizing saliva as a noninvasive sample, we identified 26 metabolites with differential expression in smokers and 31 in ex-smokers. However, no such significant alteration was observed in the non-smokers subgroup. The multivariate analysis distinctly separated the COPD subgroups from healthy controls. Additionally, pathway enrichment analysis revealed perturbations in key metabolic pathways, including unsaturated fatty acid biosynthesis, arginine biosynthesis, the tricarboxylic acid (TCA) cycle, and pyruvate metabolism. Moreover, univariate Random forest analysis identified four metabolites (cyclopentanone, tetradecane 4-methyl, acetophenone, and scyllo-inositol) as potential biomarkers distinguishing COPD subgroups from healthy controls. This study offers novel molecular insights into the association of smoking with disease progression and provides a mechanistic understanding of COPD in different subgroups for better management of the disease.