Sample barcoding-associated technical variation in probe-based single-cell RNA sequencing
Sample barcoding-associated technical variation in probe-based single-cell RNA sequencing
Weir, J. A.; Krebs, Y.; Chen, F.
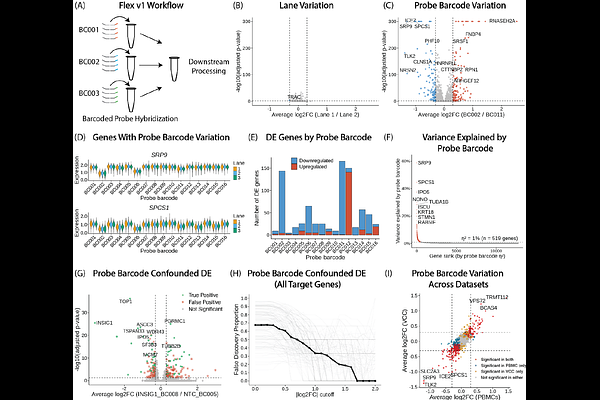
AbstractProbe-based single cell RNA sequencing approaches are increasingly becoming a technology of choice for profiling gene expression at scale and in archival tissues. The 10x Genomics Flex v1 assay enables cost-effective and high-sensitivity single-cell RNA sequencing by splitting samples across up to 16 uniquely barcoded probe sets before pooling and loading onto a single lane of a microfluidic chip. A natural consequence of this design is to leverage probe set barcoding as a sample barcoding strategy for case-control experiments. However, we observed that Flex v1 probe set barcode identity drives substantial technical variation between probe set barcodes, an effect that is reproducible across lanes and independent datasets. When Flex v1 probe set barcodes are confounded with biological sample identity, a concerning number of differentially expressed genes at standard thresholds are false positives. The Flex v2 assay, which decouples sample barcoding from probe set hybridization, significantly reduces this artifact. As the field continues to expand adoption of probe-based assays, our findings introduce probe set barcoding as an underappreciated source of technical variation in single-cell assays and emphasize the importance of experimental design when using probe-based sequencing technologies.