Temporal shifts in polygenic traits track major epidemics in Western Eurasia
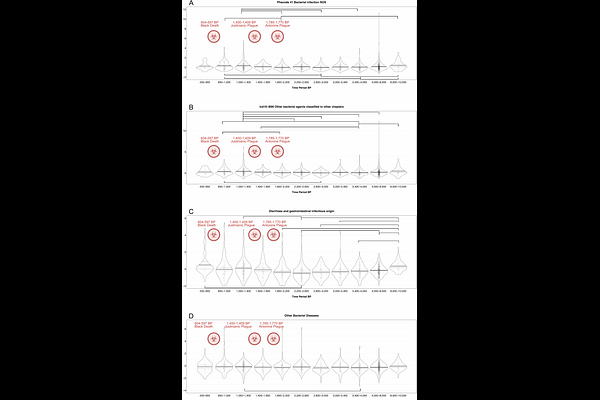
Temporal shifts in polygenic traits track major epidemics in Western Eurasia
De Angelis, F.; Fehren-Schmitz, L.; G. Amorim, C. E.
AbstractInfectious diseases are recognized as one of the strongest selective forces, exerting a profound influence on the genetic makeup of human populations over time. Recently, large-scale genome-wide association studies (GWAS) on immunological traits have underscored the notion that genetic predisposition to infectious diseases likely stems from the contribution of several thousand loci across the human genome. To model the polygenic inheritance of these traits, multiple variants can be combined into polygenic risk scores (PRS), which estimate an individual's genetic predisposition for a trait. By combining present-day GWAS statistics from large European biobanks with genomic data from more than 3,500 ancient individuals from Western Eurasia, we characterize temporal changes in four highly heritable infectious disease-related traits over a span of 10,000 years. In doing so, we account for variation in these traits across time, space, and genetic ancestries, and demonstrate that the observed patterns cannot be explained by genetic drift alone. Our findings suggest that major disease outbreaks in human history are associated with shifts in polygenic traits in human populations. Specifically, three events (the Justinian Plague, Antonine Plague, and early medieval measles outbreaks) coincide with significant shifts in the polygenic profiles of these traits. Using a Gene Ontology enrichment approach, we show that these shifts involve multiple systemic biological processes, with a consistent emphasis on metabolic pathways modulating immunological responses both directly and indirectly.