Small-molecule activators of the Staphylococcus aureus ClpC/ClpP AAA+ protease
Small-molecule activators of the Staphylococcus aureus ClpC/ClpP AAA+ protease
Jenne, T.; Viliuga, V.; Uhrig, U.; Jehle, B.; Schwan, M.; Kopp, J.; Flemming, D.; Seebach, E.; Sinning, I. M.; Bukau, B. G.; Mogk, A.
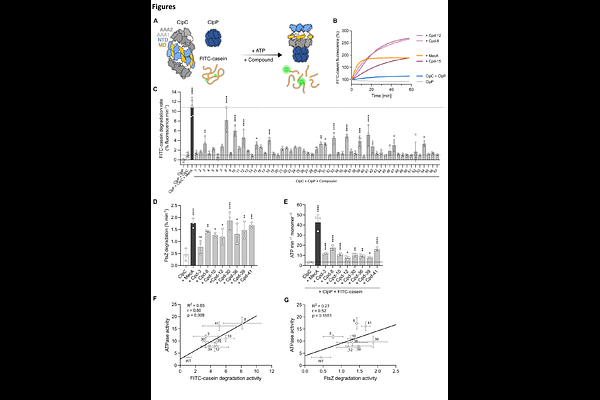
AbstractThe central AAA+ ClpC/ClpP protease in Gram-positive bacteria is crucial for virulence and stress resistance and has been recognized as drug target. Natural cyclic peptides deregulate the essential Mycobacterium tuberculosis ClpC1 and cause cell death. Similarly, overactivated mutants of the non-essential Staphylococcus aureus ClpC homologue cause uncontrolled proteolysis and severe toxicity in vivo. However, no chemical modulators of S. aureus ClpC have been described. Here, using a biochemical high-throughput screen we identify eight chemically distinct bona fide small molecules that robustly stimulate ClpC ATPase and proteolytic activity in vitro. Structural, computational, and mutational analyses define two ligandable regulatory sites within the ClpC N-terminal domain (NTD) as compound targets: a conserved hydrophobic groove and an allosteric pArg1 pocket, both engaged in substrate recognition. These findings establish S. aureus ClpC as chemically targetable and provide mechanistic insight into its regulatory architecture, enabling future development and optimization of chemical probes to deregulate AAA+ protease control.