Tracking eco-evolutionary dynamics among lung pathobiome members from children with cystic fibrosis
Tracking eco-evolutionary dynamics among lung pathobiome members from children with cystic fibrosis
Schwyter, L.; Kuratli, R.; Provveduto, S.; Do Carmo Silva, P.; Latzin, P.; Harrison, F.; Hilty, M.; Kuemmerli, R.
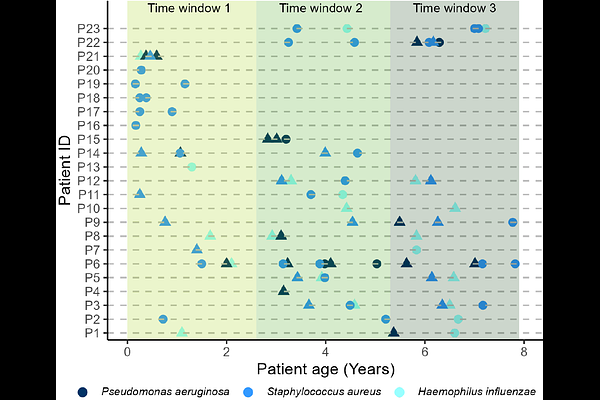
AbstractPolymicrobial lung infections are common in individuals with cystic fibrosis (CF). Pathogen communities typically follow an ecological succession in which early colonizers such as Haemophilus influenzae and Staphylococcus aureus are later joined by Pseudomonas aeruginosa. Although adaptation to the lung environment is well described for single pathogens in adult people with CF, much less is known about the role of pathogen interactions and how changes in individual pathogens influence community dynamics at the early stages of disease. To address these questions, we combined genome sequencing with phenotypic screens and pathogen interaction assays using longitudinal clinical isolates (19 P. aeruginosa, 44 S. aureus, and 21 H. influenzae) collected from 23 children with CF (0-7.9 years of age) enrolled in the SCILD (Swiss CF Infant Lung Development) cohort. Our analysis revealed that early pathogen communities are characterized by a combination of strain turnover and persistence of isolates undergoing first steps of within-host evolution. Most notably, quorum-sensing-deficient P. aeruginosa variants repeatedly emerged, showing reduced protease production and diminished inhibition of S. aureus and H. influenzae. These changes indicate that P. aeruginosa becomes less antagonistic towards co-occurring pathogens, possibly promoting community stability. Together, our results show that ecological and evolutionary dynamics between pathobiome members may play an underappreciated role in shaping CF lung disease during early childhood.