Evolutionary analysis of V protein pseudogenization in an RNA editing-deficient paramyxovirus
Evolutionary analysis of V protein pseudogenization in an RNA editing-deficient paramyxovirus
Rakib, T. M.; Akter, L.; Matsumoto, Y.
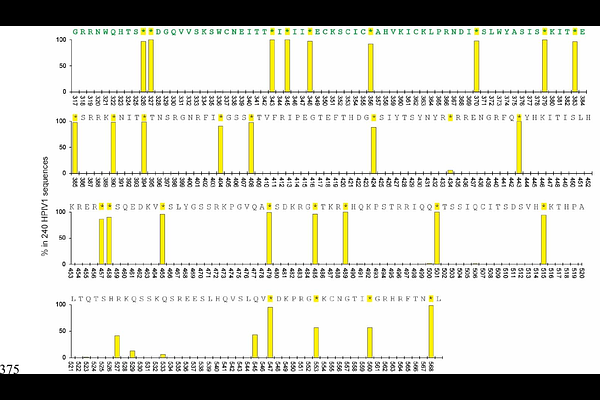
AbstractIn most paramyxoviruses, RNA editing in the P gene enables expression of the V protein. Human parainfluenza virus type 1 (HPIV-1) differs from most paramyxoviruses in that it lacks RNA editing and does not produce a functional V protein, although its genome retains sequences corresponding to the ancestral V reading frame. Here, we analyzed all HPIV-1 genome sequences available in the NCBI GenBank database to assess the evolutionary state of this V protein-specific region. Using Sendai virus (SeV) as a closely related reference with an identical P gene length, we defined a pseudo-V reading frame by virtually inserting a single nucleotide at the conserved RNA editing site. In this pseudo-V frame, HPIV-1 showed a marked excess of stop codons within the 253-amino-acid region corresponding to the post-editing sequence, far exceeding expectations under random codon usage. This pattern was not observed in other viral genes analyzed under the same definition, nor in SeV, nor was it reproduced by in silico evolutionary simulations under constraints preserving the primary open reading frame. These results are consistent with a virus-specific evolutionary trajectory following the loss of RNA editing, rather than with generic coding constraints acting on overlapping reading frames.