MiCBuS: Marker Gene Mining for Unknown Cell Types Using Bulk and Single Cell RNA-Seq Data
MiCBuS: Marker Gene Mining for Unknown Cell Types Using Bulk and Single Cell RNA-Seq Data
Zhang, S.; Lu, Y.; Luo, Q.; An, L.
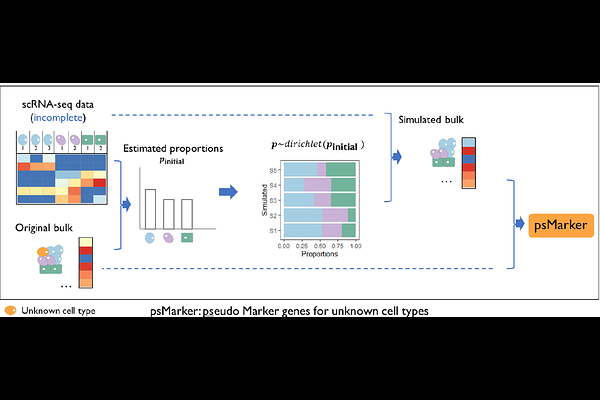
AbstractMotivation: Identifying cell type-specific expressed genes (marker genes) is essential for understanding the roles and interactions of cell populations within tissues. To achieve this, the traditional differential analysis approaches are often applied to individual cell-type bulk RNA-seq and single-cell RNA-seq data. However, real-world datasets often pose challenges, such as heterogeneous bulk RNA-seq and incomplete scRNA-seq. Heterogeneous bulk RNA-seq amalgamates gene expression profiles from multiple cell types and results in low resolution, while incomplete scRNA-seq does not capture some cell types from the tissue, leading to unknown cell types. Traditional methods fail to identify marker genes for such unknown cell types. Results: MiCBuS addresses this limitation by generating Dirichlet-pseudo-bulk RNA-seq based on bulk and incomplete single-cell RNA-seq data. By performing differential analysis of gene expressions on bulk and Dirichlet-pseudo-bulk RNA-seq samples, MiCBuS can identify the marker genes of unknown cell types, enabling the identification and characterization of these elusive cellular components. Simulation studies and real data analyses demonstrate that MiCBuS reliably and robustly identifies marker genes specific to unknown cell types, a capability that traditional differential analysis methods cannot achieve.