Full-Length Structural Modeling of Mitofusins with AlphaFold Reveals a Novel Cross-Type Dimerization and Insights into Oligomerization
Full-Length Structural Modeling of Mitofusins with AlphaFold Reveals a Novel Cross-Type Dimerization and Insights into Oligomerization
Versini, R.; Baaden, M.; Bonvin, A.; Fuchs, P.; Taly, A.
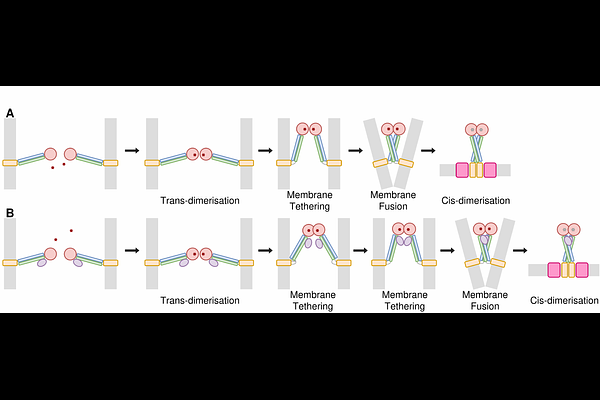
AbstractMitochondrial dynamics, involving fission and fusion, are critical for the maintenance, function, distribution, and inheritance of mitochondria, allowing their morphology to adapt to the cell's physiological needs. Mitofusins, large GTPase transmembrane proteins, play a role in this process by driving the tethering and fusion of mitochondrial outer membranes. Dysfunction in mitofusins has been associated with neurodegenerative diseases such as Parkinson's, Alzheimer's, Huntington's, and Charcot-Marie-Tooth type 2A, as well as various cancers, where their dysregulated expression influences cell proliferation, invasion, and chemotherapy resistance. Despite their importance, the precise molecular mechanisms underlying mitofusin-mediated fusion remain unclear and require further structural elucidation. In fact, no complete high-resolution structures exist for mitofusins, including their yeast homolog, Fzo1. Here, we generated and analyzed full-length structural models of mitofusins using AlphaFold. Monomeric, dimeric, and tetrameric assemblies were produced, including complexes with fusion partners Ugo1 and SLC25A46. While AlphaFold predicted limited conformational diversity for isolated monomers, structural variability emerged for predicted homo- and hetero-oligomers. Notably, our models reveal a previously undescribed cross-type dimerization mode involving interactions between heptad repeat domains, not reported in current experimental structures. Comparison with recently resolved experimental data further supports the structural relevance of this interface. These full-length models allowed us to propose a new hypothetical mechanism of outer mitochondrial membrane fusion.