dreampy: Pseudobulk mixed-model differential expression for single-cell RNA-seq in Python
Voice is AI-generated
Connected to paperThis paper is a preprint and has not been certified by peer review
dreampy: Pseudobulk mixed-model differential expression for single-cell RNA-seq in Python
Wells, S. B.; Shahnawaz, H.; Jones, J. L.
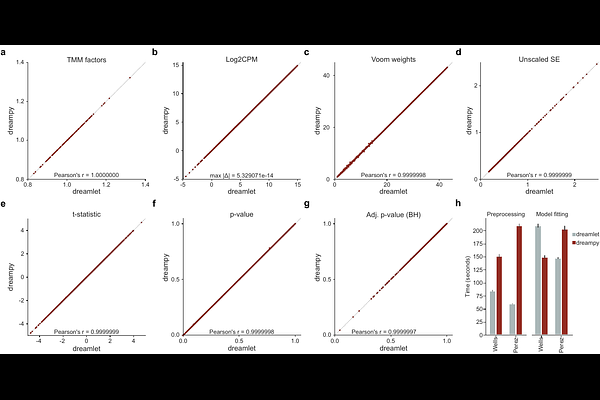
Abstractdreampy is a Python implementation of the R dreamlet framework for pseudobulk differential expression analysis of single-cell RNA-seq data. dreamlet combines voom precision-weighted linear mixed models with empirical Bayes moderation to handle batch effects, repeated measures, and other hierarchical structure in multi-donor studies, but exists entirely within the R/Bioconductor ecosystem. dreampy reproduces this pipeline natively in Python, integrating with AnnData and the scverse ecosystem.