A comprehensive benchmark of publicly available image foundation models for their usability to predict gene expression from whole slide images
A comprehensive benchmark of publicly available image foundation models for their usability to predict gene expression from whole slide images
Jabin, A.; Ahmad, S.
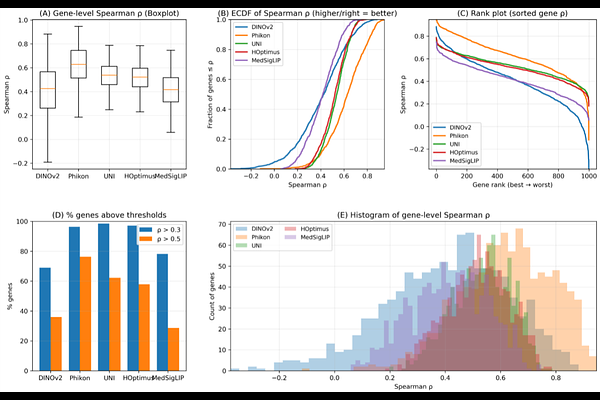
AbstractRecent advances in large-scale self-supervised learning have led to the emergence of foundation models capable of extracting transferable visual representations from high-dimensional image data. In computational pathology, such models are increasingly used as feature encoders for molecular prediction tasks. However, systematic benchmarking of publicly available image foundation models for transcriptomic prediction from whole-slide images (WSIs) remains limited. Here, we perform a comprehensive evaluation of five state-of-the-art vision foundation models-DINOv2, Phikon, UNI, H-Optimus-0, and MedSigLIP-for gene expression prediction using the TCGA-BRCA cohort. Tile embeddings extracted from each model were aggregated via attention-based multiple instance learning (MIL), followed by multi-target regression to predict RNA-seq expression profiles. Performance was assessed using gene-level Spearman correlation across samples. Histopathology-specific foundation models consistently outperformed general-purpose encoders, with Phikon achieving the strongest overall performance, followed by UNI and H-Optimus-0. These findings demonstrate that domain-aligned pretraining substantially enhances morphology-to-transcriptome inference and provide a principled benchmark for foundation model selection in molecular pathology.