Efficient and scalable Python implementation of ANCOM-BC for omics differential abundance testing
Efficient and scalable Python implementation of ANCOM-BC for omics differential abundance testing
Wu, Z.; Lin, H.; Morton, J.; Zhu, Q.
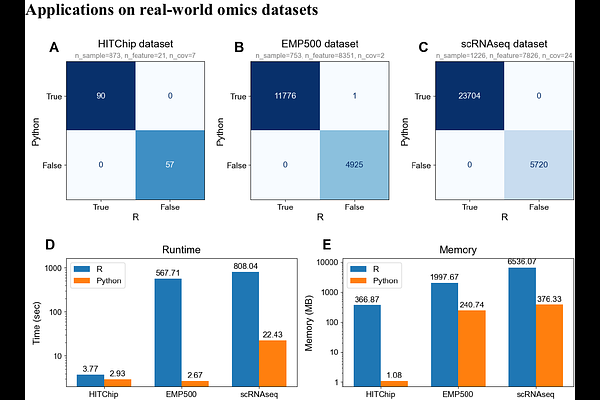
AbstractANCOM-BC (Analysis of Compositions of Microbiomes with Bias Correction) is a critical algorithm for differential abundance analysis. Designed for handling compositional data, it features the ability to estimate the unknown sampling fractions and correct the bias induced by their differences. While the original R package is robust for standard dataset sizes, the increasing magnitude of modern multi-omics studies presents computational challenges, often resulting in runtime bottleneck for high-dimensional data. Here, we present a native Python implementation of ANCOM-BC in the scikit-bio package that significantly accelerates the workflow, enabling its application to large-scale datasets. By leveraging the vectorization with NumPy and numerical optimization in SciPy, we enhanced the efficiency without compromising the statistical accuracy of the method. The benchmarking results revealed a dramatic increase in performance in both simulated and real datasets. We demonstrated that our Python implementation generates test results that match the original R package, while achieving a 100-fold acceleration in runtime and an 87% reduction in memory consumption. This new implementation allows for the analysis of datasets that are significantly larger than previously possible, and enables applications to alternative omics data types and complex statistical models that vary greatly by the number of samples, features, and covariates.