Predicting Pre-treatment Resistance or Post-treatment Effect? A Systematic Benchmarking of Single-Cell Drug Response Models
Predicting Pre-treatment Resistance or Post-treatment Effect? A Systematic Benchmarking of Single-Cell Drug Response Models
Shen, L.; Sun, X.; Zheng, S.; Hashmi, A.; Eriksson, J.; Mustonen, H.; Seppänen, H.; Shen, B.; Li, M.; Vähä-Koskela, M.; Tang, J.
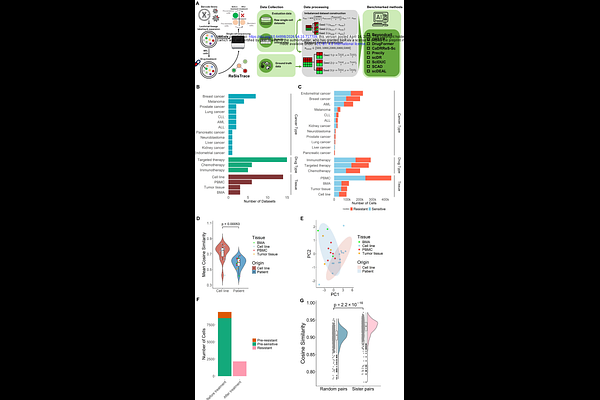
AbstractIntratumoral heterogeneity is a major driver of variable drug responses in cancer. Single-cell RNA sequencing (scRNA-seq) enables the characterization of such heterogeneity, providing an opportunity to predict drug response at single-cell resolution. As a result, a growing number of computational models have been developed to infer drug response from scRNA-seq datasets. However, their performance, robustness, and generalizability across different biological contexts have not been systematically evaluated. To address this gap, we conducted a comprehensive benchmarking of representative single-cell drug response prediction models. Using 26 curated datasets comprising over 760,000 cells across 12 cancer types and 21 therapeutic agents, we constructed balanced and imbalanced scenarios to reflect more realistic distributions of drug response labels. To address the lack of ground-truth drug-response labels in conventional scRNA-seq datasets, we further incorporated lineage-tracing data with experimentally validated drug-response annotations, enabling model evaluation in a clinically relevant pre-treatment prediction setting. Our results show that across the tested methods, the prediction performance is markedly higher in cell lines than in tissue samples. Under imbalanced conditions, most methods exhibited sharp performance declines, whereas scDEAL demonstrated the highest robustness. Independent validation using an in-house pancreatic ductal adenocarcinoma dataset further confirms the robustness of scDEAL and its ability to capture biologically meaningful state transitions. Label-substitution experiment revealed that this robust performance partially driven by the model's specific training label construction. However, the benchmarking with lineage-tracing data reveals a fundamental limitation: most models capture drug-induced transcriptional changes but struggle to predict a cell's intrinsic resistance state prior to treatment. In summary, our study not only defines the performance boundaries of current approaches but also highlights their limitations in addressing intratumoral heterogeneity, extreme class imbalance, and the prediction of intrinsic cellular resistance, emphasizing the need for the development of next-generation single-cell drug response models with stronger clinical relevance.