IMAS enables target-aware integration of tumour multiomics to resolve communication-guided regulatory mechanisms
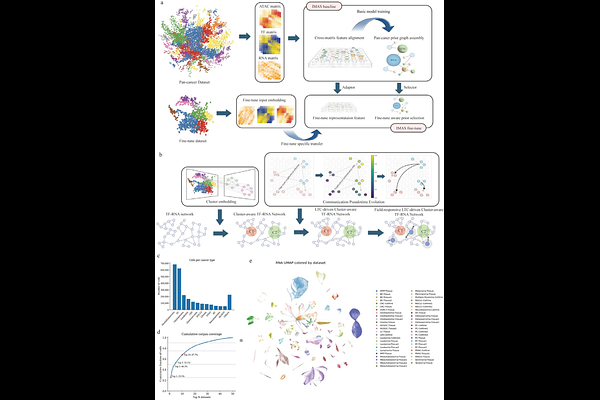
IMAS enables target-aware integration of tumour multiomics to resolve communication-guided regulatory mechanisms
Deyang, W.; Yamashiro, T.; Inubushi, T.
AbstractTumour multiomic datasets are often sparse, heterogeneous and limited in size, hindering robust and interpretable discovery of regulatory mechanisms. Here we present IMAS, a target-aware integrative framework for multiomic data augmentation and mechanism prioritization that leverages a pan-cancer single-cell multiomic resource to contextualize new tumour datasets and identify reliable sample-specific mechanistic hypotheses. IMAS combines shared latent-space modelling with target-domain adaptation to improve correspondence between predicted and observed RNA and TF profiles while concentrating explanatory predictive supports within the target dataset. Building on this adapted representation, IMAS reconstructs structured RNA-TF coupling networks, refines intercellular signaling through ligand-informed communication modelling, and organizes regulatory programs along communication-associated ordering. In independent colon cancer data, IMAS improved cluster-resolved correspondence and revealed communication-guided regulatory cascades across malignant epithelial states. A LAMB1-centred analysis further demonstrates how the framework supports progressive reinforcement of local regulatory structure and enables perturbation-based probing of context-specific dependencies. Rather than exhaustively predicting all possible outcomes, IMAS provides a target-aware and interpretable strategy to construct consistent and interpretable mechanism-discovery scaffolds and prioritize regulatory dependencies in data-limited tumour systems.