Revisiting Reconstruction Likelihood: Variational Autoencoders for Biological and Biomedical Data Clustering
Revisiting Reconstruction Likelihood: Variational Autoencoders for Biological and Biomedical Data Clustering
Korenic, A.; Özkaya, U.; Capar, A.
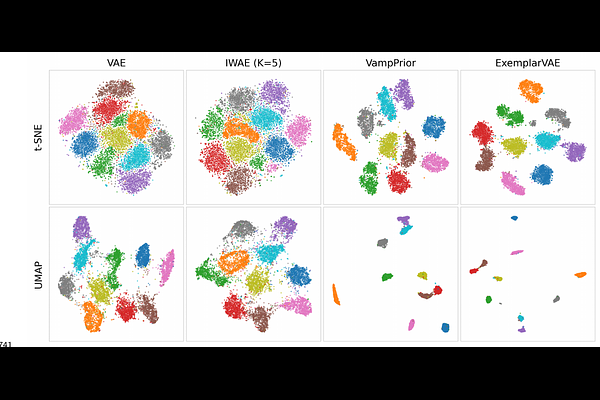
AbstractBackground and Objective: Variational Autoencoders (VAEs) offer a powerful framework for unsupervised anomaly detection and data clustering, often surpassing traditional methods. A core strength of VAEs lies in their ability to model data distributions probabilistically, enabling robust identification of anomalies and clusters through reconstruction likelihood --- a stochastic metric providing a principled alternative to deterministic error scores. Methods: We investigated how different VAE architectures, combining reconstruction likelihood with a learnable or data-driven prior, performed in a clustering task on a toy dataset such as MNIST. Results were verified using dimensionality reduction techniques like t-distributed Stochastic Neighbor Embedding (t-SNE) and Uniform Manifold Approximation and Projection (UMAP), alongside clustering algorithms such as k-means and Hierarchical Density-Based Spatial Clustering of Applications with Noise (HDBSCAN). Results: The VAE's encoder inherently maps data points into a latent space exhibiting discernible cluster structure, as evidenced by alignment with ground truth labels. While dimensionality reduction techniques (both t-SNE and UMAP) facilitated the application of clustering algorithms (k-means and HDBSCAN), these methods were primarily used to visualize and interpret the latent space organization. Conclusions: This study demonstrates that VAEs effectively cluster data by implicitly encoding assignments in their latent representations. Determining cluster membership from encoder output, combined with reconstruction likelihood using semantic features, offers a principled approach for identifying typical samples and anomalies. Future research should focus on leveraging this inherent clustering capability of VAEs to enhance interpretability and facilitate clinical application.