Rastair: an integrated variant and methylation caller
Rastair: an integrated variant and methylation caller
Etzioni, Z.; Zhao, L.; Hertleif, P.; Schuster-Boeckler, B.
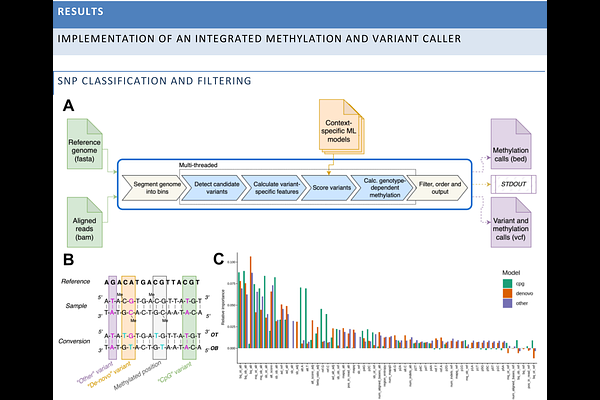
AbstractCytosine methylation is a crucial epigenetic mark that impact tissue-specific chromatin conformation and gene expression. For many years, bisulfite sequencing (BS-seq), which converts all non-methylated cytosine (C) to thymine (T), remained the only approach to measure cytosine methylation at base resolution. Recently, however, several new methods that convert only methylated cytosines to thymine (mC[->]T) have become widely available. Here we present rastair, an integrated software toolkit for simultaneous SNP detection and methylation calling from mC[->]T sequencing data such as those created with Watchmaker's TAPS+ and Illumina's 5-Base chemistries. Rastair combines machine-learning-based variant detection with genotype-aware methylation estimation. Using NA12878 benchmark datasets, we show that rastair outperforms existing methylation-aware SNP callers and achieves F1 scores exceeding 0.99 for datasets above 30x depth, matching the accuracy of state-of-the-art tools run on whole-genome sequencing data. At the same time, rastair is significantly faster than other genetic variant callers, processing a 30x depth file takes less than 30 minutes given 32 CPU cores on an Intel Xeon, and half as long when a GPU is available. By integrating genotyping with methylation calling, rastair reports an additional 500,000 positions in NA12878 where a SNP turns a non-CpG reference position into a "de-novo" CpG. Vice-versa, rastair also identifies positions where a variant disrupts a CpG and corrects their reported methylation levels. Rastair produces standard-compliant outputs in vcf, bam and bed formats, facilitating integration into downstream analyses pipelines. Rastair is open-source and available via conda, Dockerhub, and as pre-compiled binaries from https://www.rastair.com.